A good preprocessing pipeline can help in training models with higher accuracy by feeding clean data.
ESR4 is an integral position in CLARIFY Project forming connection among the researchers from Blockchain technology and Artificial Intelligence. My research project is designed for several outcomes such as preprocessing methods and algorithms. In computer vision, processing an image requires several computational methods to evaluate different statistical figures. It plays a vital role in understanding the image at a micro-level.
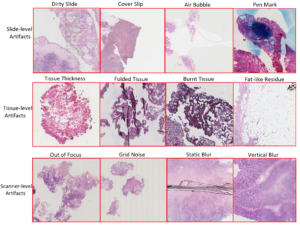
Research in digital pathology (DP) is observing an increase in the use of Machine Learning algorithms. Training deep learning models with histopathological data requires digital pathology expertise and understanding to annotate whole slide images (WSI). Deep Learning (DL) experts are not familiar with DP expertise and vice versa. It is crucial to label regions of interest to deliver good feature level information to the model. The WSIs are usually related to metadata and clinical notes. The microdata in WSI is very huge and contains a lot of artefacts. These abnormalities can be broadly categorized into three streams 1) Slide-level artefacts, 2) Tissue level artefacts and 3) Scanner-level artefacts. These artefacts result from improper fixation, freezing, impregnation and paraffin wax. The figure below demonstrates H&E patches with different artefacts.
Combining traditional methods with deep learning frameworks can obtain a progressively high performing Computer-Aided Design (CAD) system. Training DL models with a bigger dataset of high-resolution images is a limitation for hardware resources. Deep learning frameworks are powerful in analysing complex patterns from gigapixel images. Computational pathology urges the need for efficient algorithms for classification, segmentation, and detection. Deep learning researchers spend most of their time understanding the DP data and creating an automatic method to omit these artefacts.
A preprocessing pipeline can be designed to cover all necessary filters to eliminate images that may deteriorate model performance. The selection of image filters and their thresholds plays a significant role in their computation. This parametric assessment may be tuned for different dataset based on lightness, cracks, pen marks, tissue fats and other tissue anomalies. Before proceeding with this sequence of filters image, quality checks are necessary. Image quality can be determined based on basic information such as magnification, format, and other derived information such as blur ratio and pixel-to-use. The grid patching method is suitable to perform image analysis in a further step. Stain normalization is the most popular step in preprocessing. In the light of its significance in digital pathology, we propose various techniques to deal with colour variability and their time complexity. Lastly, patch selection is useful to make an unbiased classifier or removing correlation from other parts of tissue types in this pipeline.
Neel Kanwal – ESR4